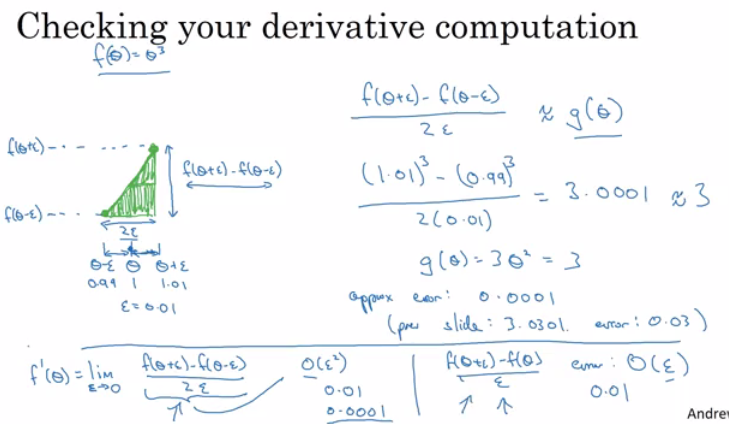
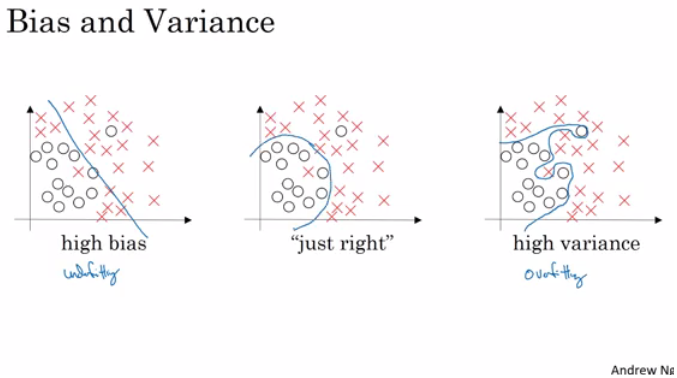
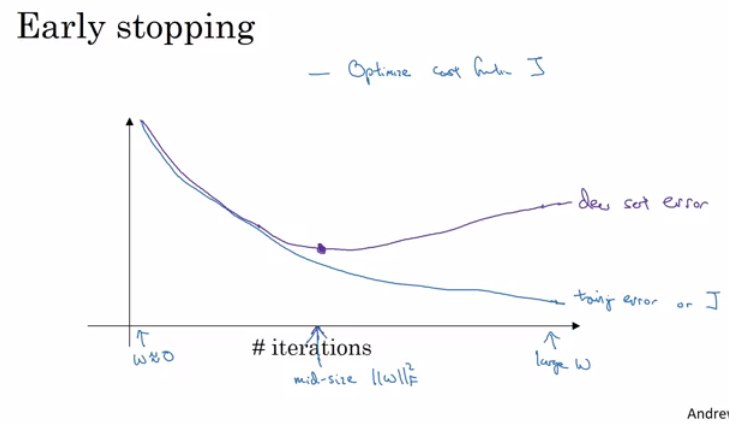
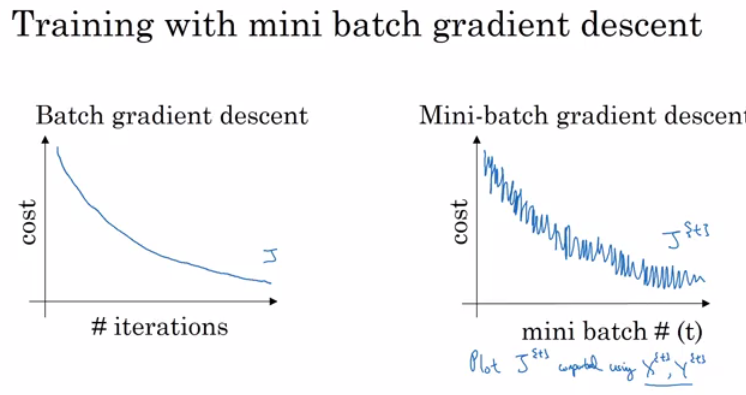
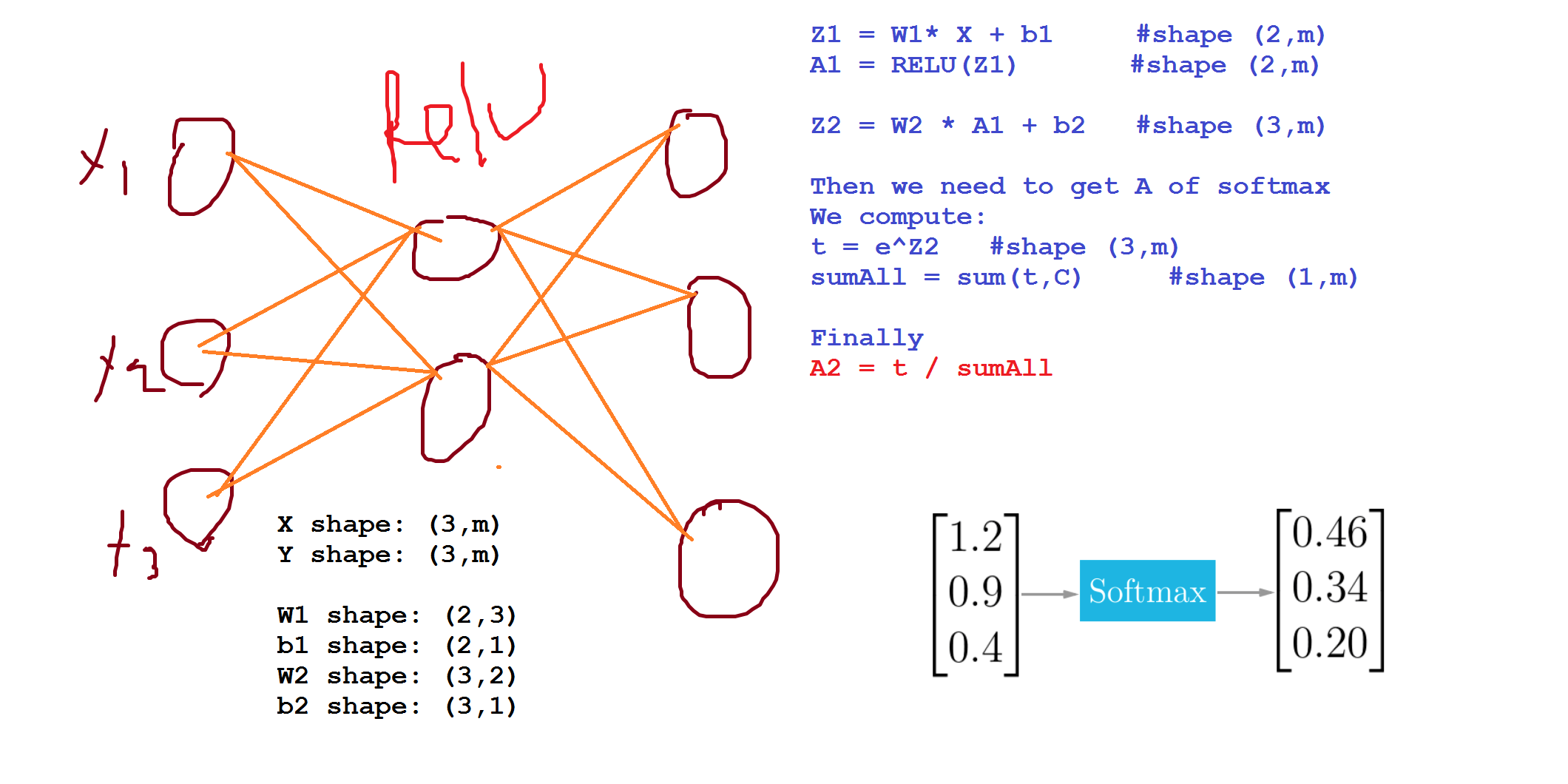
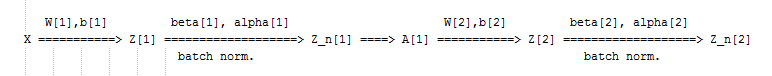Rather than the deep learning process being a black box, you will understand what drives performance, and be able to more systematically get good results. You will also learn TensorFlow.
1
2
3
4
5
6
| v = 0
Repeat
{
Get theta(t)
v = beta * v + (1-beta) * theta(t)
}
|
1
| v(t) = (beta * v(t-1) + (1-beta) * theta(t)) / (1 - beta^t)
|
1
2
3
4
5
6
7
8
9
| vdW = 0, vdb = 0
on iteration t:
# can be mini-batch or batch gradient descent
compute dw, db on current mini-batch
vdW = beta * vdW + (1 - beta) * dW
vdb = beta * vdb + (1 - beta) * db
W = W - learning_rate * vdW
b = b - learning_rate * vdb
|
1
2
3
4
5
6
7
8
9
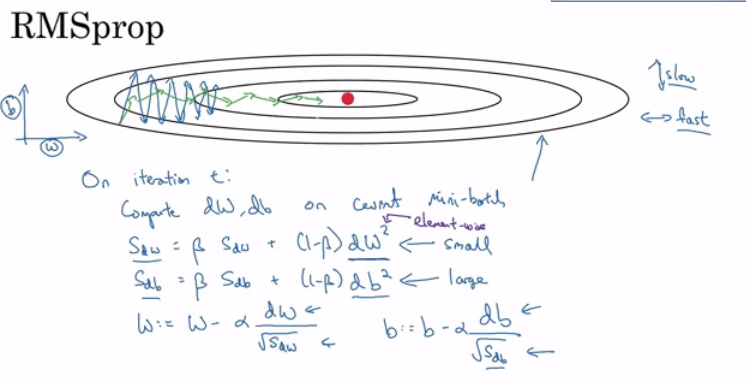
| sdW = 0, sdb = 0
on iteration t:
# can be mini-batch or batch gradient descent
compute dw, db on current mini-batch
sdW = (beta * sdW) + (1 - beta) * dW^2 # squaring is element-wise
sdb = (beta * sdb) + (1 - beta) * db^2 # squaring is element-wise
W = W - learning_rate * dW / sqrt(sdW)
b = B - learning_rate * db / sqrt(sdb)
|
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
| vdW = 0, vdW = 0
sdW = 0, sdb = 0
on iteration t:
# can be mini-batch or batch gradient descent
compute dw, db on current mini-batch
vdW = (beta1 * vdW) + (1 - beta1) * dW # momentum
vdb = (beta1 * vdb) + (1 - beta1) * db # momentum
sdW = (beta2 * sdW) + (1 - beta2) * dW^2 # RMSprop
sdb = (beta2 * sdb) + (1 - beta2) * db^2 # RMSprop
vdW = vdW / (1 - beta1^t) # fixing bias
vdb = vdb / (1 - beta1^t) # fixing bias
sdW = sdW / (1 - beta2^t) # fixing bias
sdb = sdb / (1 - beta2^t) # fixing bias
W = W - learning_rate * vdW / (sqrt(sdW) + epsilon)
b = B - learning_rate * vdb / (sqrt(sdb) + epsilon)
|
1
2
3
| Z[l] = W[l]A[l-1] + b[l] => Z[l] = W[l]A[l-1]
Z_norm[l] = ...
Z_tilde[l] = gamma[l] * Z_norm[l] + beta[l]
|
1
2
| t = e^(Z[L]) # shape(C, m)
A[L] = e^(Z[L]) / sum(t) # shape(C, m), sum(t) - sum of t's for each example (shape (1, m))
|
1
| L(y, y_hat) = - sum(y[j] * log(y_hat[j])) # j = 0 to C-1
|
1
| J(w[1], b[1], ...) = - 1 / m * (sum(L(y[i], y_hat[i]))) # i = 0 to m
|
1
2
3
| W1 = tf.get_variable("W1", [25,12288], initializer = tf.contrib.layers.xavier_initializer(seed = 1))
b1 = tf.get_variable("b1", [25,1], initializer = tf.zeros_initializer())
|